Data analysis and uncertainty evaluation
The aim of this work package is to improve the current state of the art in data analysis for perfusion imaging of the myocardium. Through careful study of images from the phantom being developed in WP1, partners will thoroughly assess current deconvolution methods and develop alternatives using both parametric and non parametric approaches.
WP2 begins with a metrological assessment of existing deconvolution methods and is followed by uncertainty characterisation for deconvolution. Bayesian methodologies and extensions to existing deconvolution techniques are then introduced. Classification techniques will be investigated to identify disease-related high and low perfusion states. Finally CMR perfusion results will be compared with those from other imaging modalities such as PET to assess consistency for measurements of the phantom. Key to this activity will be the reliable measurement uncertainties that will be established in this work package.
Validation of the perfusion quantification process, using a traceable test object
The following diagram shows a schematic of the validation of a perfusion quantification process using contrast-enhanced cardiovascular magnetic resonance imaging (CMR), from obtaining the scans to comparing the estimated flow (perfusion) to measured flow:
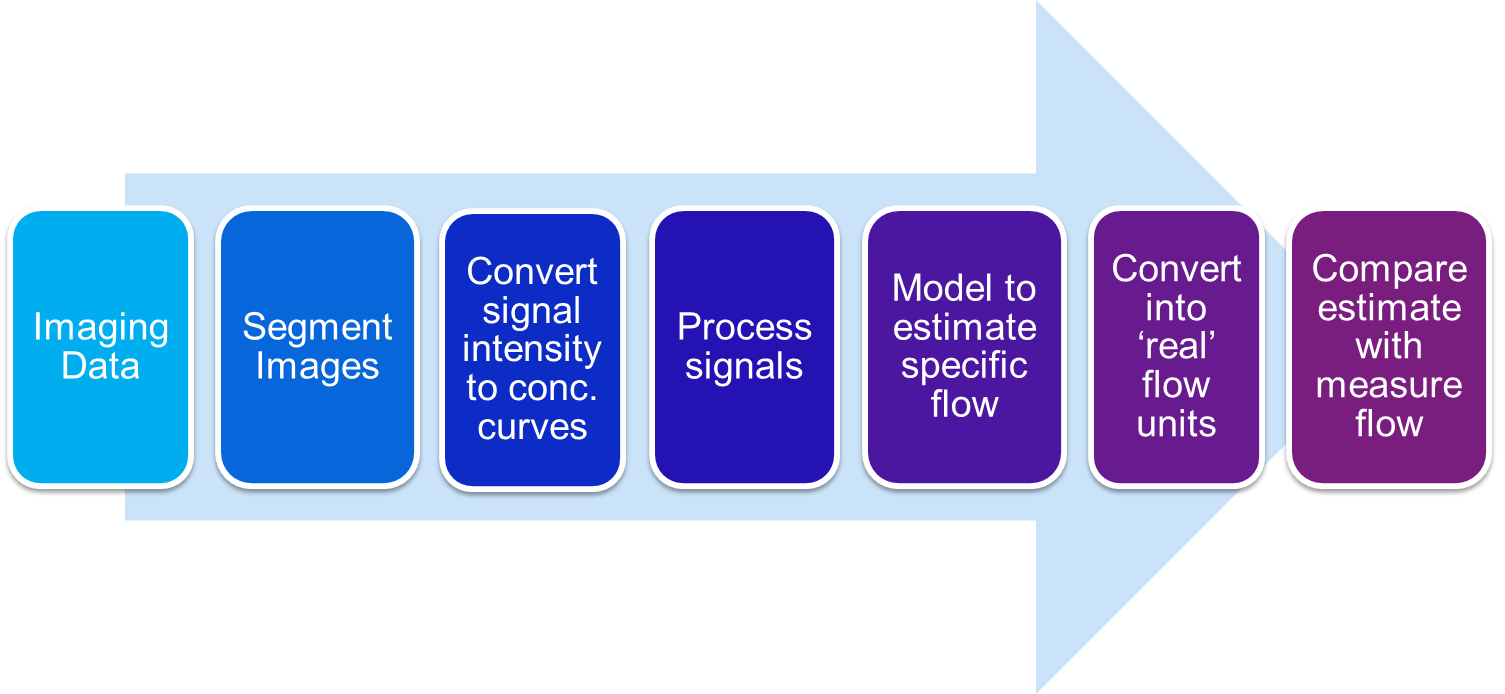
Within the phantom, each step of the process is shown below:

The comparison between estimated and measured flow can be performed using the ground-truth flow maps measured using MR flow technique. Software has been developed for pixel-wise quantification of perfusion within the phantom, as well as the validation process.
Software for pixel-wise perfusion quantification within the phantom
Software is being developed to perform the data analysis of the full pipeline of a pixel-wise perfusion quantification within the phantom, including key steps such as:



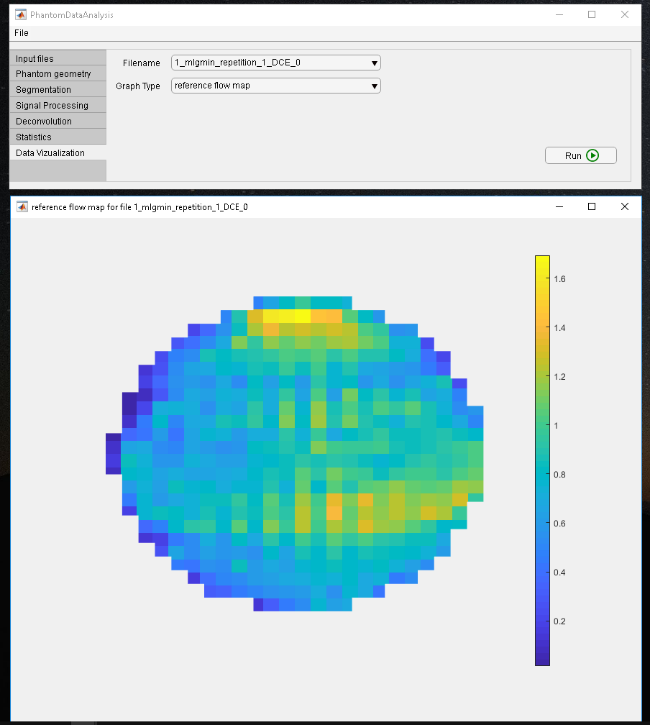
Pixel-wise quantification of myocardial perfusion using spatial Tikhonov regularization
Currently, the data used for the pixel-wise analysis of cardiac perfusion are either filtered prior to a fitting procedure, which inherently reduces the spatial resolution of data; or all pixels are considered without any regularization or prior filtering, which yields an unstable fit in the presence of low signal-to-noise ratio. Within the project we have proposed a new pixel-wise analysis based on spatial Tikhonov regularization which exploits the spatial smoothness of the data and ensures accurate quantification even for images with low signal-to-noise ratio (see Lehnert et al 2018 Phys. Med. Biol. in press https://doi.org/10.1088/1361-6560/aae758).

The figure above shows a schematic of different methods in the quantification of myocardial perfusion. (a) Segment-wise analysis method: averaged myocardial signal intensities are fitted in each segment. (b) Standard pixel-wise analysis: signal intensities are fitted independently in each pixel. Temporal and/or spatial filtering can be used before the fitting. (c) Proposed method: pixel-wise quantification of unfiltered data. Linearized fitting and Tikhonov regularization are carried out simultaneously. Beforehand, an initial parameter guess is obtained by fitting the SVD-approximated curves in each pixel
We study the performance of our method on a numerical phantom and demonstrate that the method reduces significantly the root-mean square error in the perfusion estimate compared to a non-regularized fit. In patient data our method allows us to recover the myocardial perfusion and to distinguish between healthy and ischemic regions. The below figure shows the results of applying the method to five different patients and shows the corresponding perfusion maps and time delay maps. The perfusion defects are indicated by red arrows.
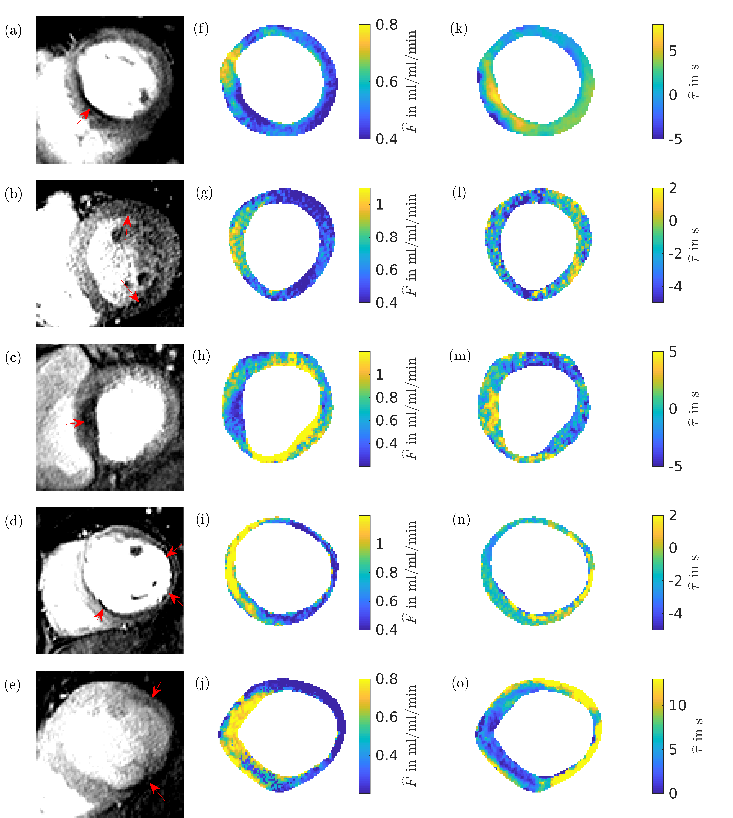
In Lehnert et al 2018 Phys. Med. Biol. in press https://doi.org/10.1088/1361-6560/aae758 we demonstrated pixel-wise quantification of myocardial perfusion by means of a spatial Tikhonov regularization in both simulated and patient data. The challenge of pixel-wise quantification is the typically low SNR. Because of this, most work on pixel-wise quantification of perfusion relies on filtering spatially and/or temporally prior to the deconvolution step. In contrast to prefiltering in time and space, the Tikhonov regularization suggested here performs fitting and regularization in one step and, therefore, balances the effects of these two procedures. Whilst errors that are introduced during the filtering that takes place before the fitting will necessarily affect the results, this is not necessarily the case for our method. Instead, the result of the Tikhonov regularization is a balance between the best fit to the (noisy) time curves and the spatial smoothness of the resulting parameters. Compared to methods which forgo any filtering or regularization, the Tikhonov regularization takes into account the spatial smoothness of the parameter as additional information, and this can reduce the errors in the parameter estimation as we were able to show for the numerical phantom.
Risk-based decisions about the degree of severity of myocardial perfusion in the presence of measurement uncertainty
A decision making framework on the degree of severity of myocardial perfusion in patients is being developed. The desired outputs of this work are:
- Classification of each patient as ischemic/non-ischemic along with an associated probability which reflects the uncertainty in the process and in the myocardial perfusion (blood flow) values
- Location of ischemia perhaps in terms of affected segments
By combining patient data and expert insights, this tool could (i) help less experienced clinicians make better decisions regarding patient health, (ii) serve as a starting point for further investigation, (iii) be used as a screening to categorise patients in order of severity, so that the most severe cases can be prioritised on the clinical list.
The following decision rule is used: if more than 10% of the pixels have a perfusion value less than 1.5 ml/g/min, then the patient is considered to have ischemia.
The figure below shows an example of a Cartesian representation of the perfusion values (ml/g/min) resulting from the analysis of perfusion PET scans for a specific patient for affected pixels (those below perfusion threshold of 1.5 ml/g/min):
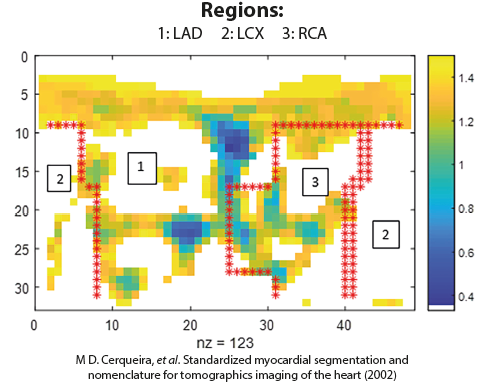
There is a need for incorporating measurement uncertainty of the perfusion maps into the decision process. In the figure above it can be seen that there are several pixels where the perfusion value is close to the threshold value, 1.5 ml/g/min. Errors in the estimated perfusion values of these pixels (and pixels with perfusion values slightly larger than 1.5 ml/g/min) could alter the diagnosis. Taking into account the uncertainty would give us a probability associated with a decision/diagnosis. Hence we can also quantify the risk to the patient or medical practice resulting from incorrect diagnosis.
Region-wise simulation was performed per-patient based on perfusion maps resulting from the analysis of PET data. The risk was evaluated for a range of uncertainty values and degrees of correlation, giving an indication of a critical value of uncertainty that would cause a change in the decision of the clinician. The results of the Monte Carlo simulations carried out are shown in the figure below:
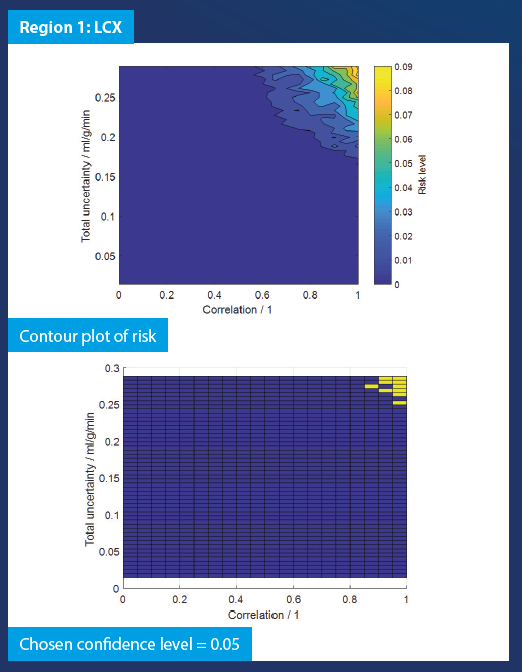
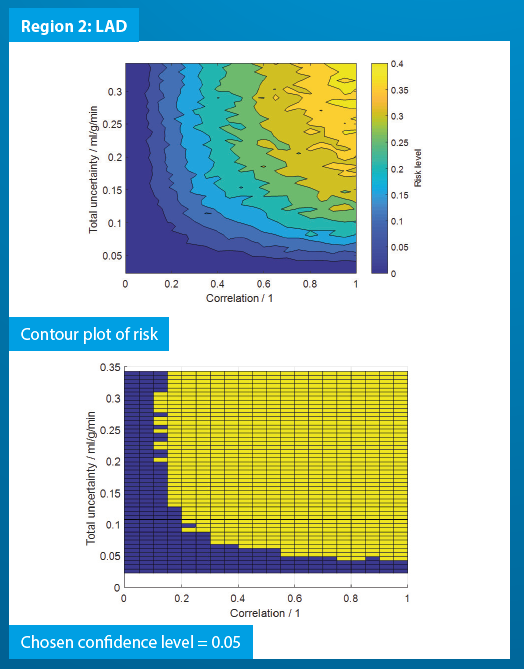


From the graphs above, we can ascertain the risks corresponding to a particular level of uncertainty and correlation. When the uncertainty and correlation are sufficiently large, there is a possibility of incorrect diagnosis of a patient. The risk levels are patient specific.

